summary method for class "nmf". Describes metadata representation in NMF factors. Returns object of class nmfSummary. Plot results using plot.
Arguments
- object
an object of class "
nmf", usually, a result of a call tonmf- group_by
a discrete factor giving groupings for samples or features. Must be of the same length as number of samples in
object$hor number of features inobject$w.- stat
either
sum(sum of factor weights falling within each group), ormean(mean factor weight falling within each group).- ...
Additional arguments (unused).
- x
nmfSummaryobject, the result of callingsummaryon annmfobject
Value
data.frame with columns group, factor, and stat
A ggplot2 object showing factor representation by group.
Examples
# \donttest{
library(Matrix)
A <- rsparsematrix(100, 50, 0.1)
model <- nmf(A, k = 3, seed = 42, tol = 1e-2, maxit = 10)
groups <- factor(sample(c("A", "B"), 50, replace = TRUE))
s <- summary(model, group_by = groups)
# }
# \donttest{
library(Matrix)
A <- rsparsematrix(100, 50, 0.1)
model <- nmf(A, k = 3, seed = 42, tol = 1e-2, maxit = 10)
groups <- factor(sample(c("A", "B"), 50, replace = TRUE))
s <- summary(model, group_by = groups)
if (requireNamespace("ggplot2", quietly = TRUE)) {
plot(s)
}
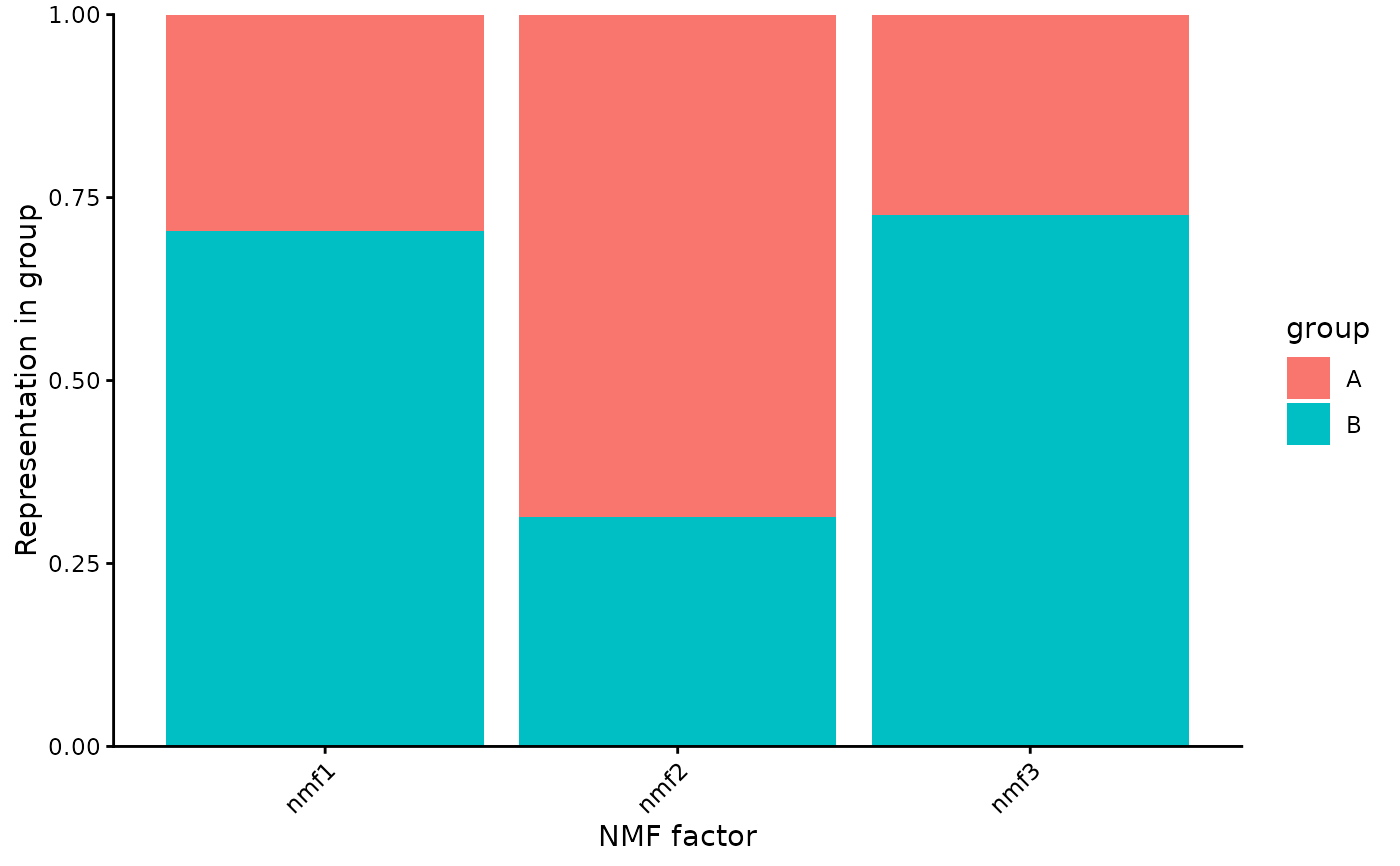 # }
# }