High-performance NMF of the form \(A = wdh\) for large dense or sparse matrices. Supports single-rank fitting or cross-validation across multiple ranks. Returns an nmf object or nmfCrossValidate data.frame.
Usage
nmf(
data,
k,
tol = 1e-04,
maxit = 100,
L1 = c(0, 0),
L2 = c(0, 0),
seed = NULL,
mask = NULL,
loss = c("mse", "gp", "nb", "gamma", "inverse_gaussian", "tweedie"),
nonneg = c(TRUE, TRUE),
test_fraction = 0,
verbose = FALSE,
projective = FALSE,
symmetric = FALSE,
zi = c("none", "row", "col"),
robust = FALSE,
...
)Arguments
- data
dense or sparse matrix of features in rows and samples in columns. Prefer
matrixorMatrix::dgCMatrix, respectively. Also accepts a file path (character string) which will be auto-loaded based on extension.- k
rank (integer), vector of ranks for cross-validation, or "auto" for automatic rank determination.
- tol
tolerance of the fit (default 1e-4)
- maxit
maximum number of fitting iterations (default 100)
- L1
LASSO penalties in the range (0, 1], single value or array of length two for
c(w, h)- L2
Ridge penalties greater than zero, single value or array of length two for
c(w, h)- seed
initialization control. Accepts:
NULLfor random init, an integer for reproducible random init, a matrix (m x k) for custom W initialization, a list of matrices for multi-init, or a string:"lanczos","irlba","randomized", or"svd"(auto-select) for SVD-based initialization.- mask
missing data mask. Accepts:
NULL(no masking),"zeros"(mask zeros),"NA"(mask NAs), a dgCMatrix/matrix (custom mask), orlist("zeros", <matrix>)to mask zeros and use a custom mask simultaneously.- loss
loss function:
"mse"(default),"gp","nb","gamma","inverse_gaussian", or"tweedie". For robust loss (Huber/MAE), use therobustparameter instead.- nonneg
logical vector of length 2 for
c(w, h)specifying non-negativity constraints (defaultc(TRUE, TRUE)).- test_fraction
fraction of entries to hold out for cross-validation (default 0 = disabled).
- verbose
print progress information during fitting (default FALSE)
- projective
if
TRUE, use projective NMF (defaultFALSE).- symmetric
if
TRUE, use symmetric NMF where H = W^T (defaultFALSE).- zi
zero-inflation mode:
"none"(default),"row", or"col". Requiresloss="gp"orloss="nb".- robust
robustness control.
FALSE(default),TRUE(Huber delta=1.345),"mae"(near-MAE via very small Huber delta), or a positive numeric Huber delta. Huber loss is quadratic for small residuals and linear for large ones, controlled by delta.TRUE(delta=1.345) provides moderate outlier robustness. A large delta (e.g. 100) approaches standard MSE; a small delta (e.g. 0.01) approaches MAE."mae"is shorthand for delta=1e-4 (effectively L1 loss).- ...
advanced parameters. See Advanced Parameters section.
Value
When k is a single integer: an S4 object of class nmf with slots:
wfeature factor matrix,
m x kdscaling diagonal vector of length
k(descending order after sorting)hsample factor matrix,
k x nmisclist containing
tol(final tolerance),iter(iteration count),loss(final loss value),loss_type(loss function used), andruntime(seconds).
When k is a vector: a data.frame of class nmfCrossValidate with columns
k, rep, train_loss, test_loss, and best_iter.
Details
This fast NMF implementation decomposes a matrix \(A\) into lower-rank non-negative matrices \(w\) and \(h\), with columns of \(w\) and rows of \(h\) scaled to sum to 1 via multiplication by a diagonal, \(d\): $$A = wdh$$
The scaling diagonal ensures convex L1 regularization, consistent factor scalings regardless of random initialization, and model symmetry in factorizations of symmetric matrices.
The factorization model is randomly initialized. \(w\) and \(h\) are updated by alternating least squares.
RcppML achieves high performance using the Eigen C++ linear algebra library, OpenMP parallelization, a dedicated Rcpp sparse matrix class, and fast sequential coordinate descent non-negative least squares initialized by Cholesky least squares solutions.
Sparse optimization is automatically applied if the input matrix A is a sparse matrix (i.e. Matrix::dgCMatrix). There are also specialized back-ends for symmetric, rank-1, and rank-2 factorizations.
L1 penalization can be used for increasing the sparsity of factors and assisting interpretability. Penalty values should range from 0 to 1, where 1 gives complete sparsity.
Set options(RcppML.verbose = TRUE) to print model tolerances to the console after each iteration.
Parallelization is applied with OpenMP using the number of threads in getOption("RcppML.threads") and set by option(RcppML.threads = 0), for example. 0 corresponds to all threads, let OpenMP decide.
Cross-Validation
When k is a vector, the function performs cross-validation to find the optimal rank:
A fraction (
test_fraction) of entries are held out for validationModels are fit for each rank in
kusing only training dataTest loss is computed on held-out entries
Returns a data.frame with k, rep, train_mse, test_mse, and best_iter for each rank/replicate
Use
plot()on the result to visualize loss vs rank
Loss Functions
The loss parameter controls the objective function:
"mse": Mean Squared Error (default). Standard Frobenius norm minimization."gp": Generalized Poisson. For overdispersed count data (e.g., scRNA-seq). Usedispersionto control per-row or global overdispersion estimation. Withdispersion="none", equivalent to KL divergence (Poisson model)."nb": Negative Binomial. Quadratic variance-mean relationship. Standard for scRNA-seq data with overdispersion. Size parameter (r) estimated per-row or globally."gamma": Gamma distribution. Variance proportional to mu^2. For positive continuous data. Dispersion (phi) estimated via MoM Pearson estimator."inverse_gaussian": Inverse Gaussian. Variance proportional to mu^3. For positive continuous data with heavy right tails."tweedie": Tweedie distribution. Variance proportional to mu^p where p is set viatweedie_power(default 1.5). Interpolates between Poisson (p=1), Gamma (p=2), and Inverse Gaussian (p=3).
For robust fitting (equivalent to Huber or MAE loss), use the robust parameter
instead of a separate loss function. Setting robust = TRUE applies Huber-weighted
IRLS with delta=1.345; robust = "mae" approximates mean absolute error.
Non-MSE losses use Iteratively Reweighted Least Squares (IRLS) which may be slower but provides better fits for count data (GP, NB) and positive continuous data (Gamma, Inverse Gaussian).
Advanced Parameters (via ...)
The following parameters can be passed via ...:
Regularization:
L21Group sparsity penalty, single value or
c(w, h)(defaultc(0,0))angularAngular decorrelation penalty,
c(w, h)(defaultc(0,0))upper_boundBox constraint on factors,
c(w, h)(defaultc(0,0)= no bound)graph_W,graph_HSparse graph Laplacian matrices for feature/sample regularization
graph_lambdaGraph regularization strength,
c(w, h)(defaultc(0,0))target_HTarget matrix (k x n) for H-side regularization. Steers H toward a desired structure during fitting. See
compute_target.target_lambdaTarget regularization strength, single value or
c(w, h)(default 0). Positive values attract H toward the target (label enrichment); negative values use PROJ_ADV eigenvalue-projected adversarial removal to suppress target-correlated structure (batch removal). Seevignette("guided-nmf")for details.
Distribution Tuning:
dispersionDispersion mode for GP loss:
"per_row","per_col","global","none"theta_init,theta_max,theta_minGP theta bounds
nb_size_init,nb_size_max,nb_size_minNB dispersion bounds
gamma_phi_init,gamma_phi_max,gamma_phi_minGamma/IG dispersion bounds
tweedie_powerTweedie variance power (default 1.5)
irls_max_iter,irls_tolIRLS convergence parameters
Zero-Inflation:
zi_em_itersEM iterations per NMF step (default 1)
Solver:
solverNNLS solver:
"auto","cholesky","cd"cd_tolCD convergence tolerance (default 1e-8)
cd_maxitCD max iterations (default 100)
h_initCustom initial H matrix
Cross-Validation:
cv_seedSeparate seed(s) for CV holdout pattern
patienceEarly stopping patience (default 5)
cv_k_rangeAuto-rank search range (default
c(2, 50))track_train_lossTrack training loss in CV (default TRUE)
Resources & Output:
threadsOpenMP threads (default 0 = all)
resourceCompute backend:
"auto","cpu","gpu"normFactor normalization:
"L1","L2","none"sort_modelSort factors by diagonal (default TRUE)
Streaming:
streamingSPZ streaming mode:
"auto",TRUE,FALSEpanel_colsPanel size for streaming (default 0 = auto)
dispatchStreamPress dispatch override
Callbacks:
on_iterationPer-iteration callback function
profileEnable timing profiling (default FALSE)
Compute Resources (CPU vs GPU)
By default (resource = "auto"), RcppML auto-detects available hardware.
All features are fully supported on CPU (the default backend).
GPU acceleration (when compiled with CUDA support) accelerates sparse and dense NMF.
GPU is experimental and falls back to CPU automatically if unavailable.
Methods
S4 methods available for the nmf class:
predict: project an NMF model (or partial model) onto new samplesevaluate: calculate mean squared error loss of an NMF modelsummary:data.framegivingfractional,total, ormeanrepresentation of factors in samples or features grouped by some criteriaalign: find an ordering of factors in onenmfmodel that best matches those in anothernmfmodelprod: compute the dense approximation of input datasparsity: compute the sparsity of each factor in \(w\) and \(h\)subset: subset, reorder, select, or extract factors (same as[)generics such as
dim,dimnames,t,show,head
References
DeBruine, ZJ, Melcher, K, and Triche, TJ. (2021). "High-performance non-negative matrix factorization for large single-cell data." BioRXiv.
Examples
# \donttest{
# basic NMF
model <- nmf(Matrix::rsparsematrix(1000, 100, 0.1), k = 10)
# cross-validation to find optimal rank
sim <- simulateNMF(200, 80, k = 5, noise = 3.0, seed = 42)
cv <- nmf(sim$A, k = 2:10, test_fraction = 0.05, cv_seed = 1:3,
tol = 1e-5, maxit = 200)
plot(cv)
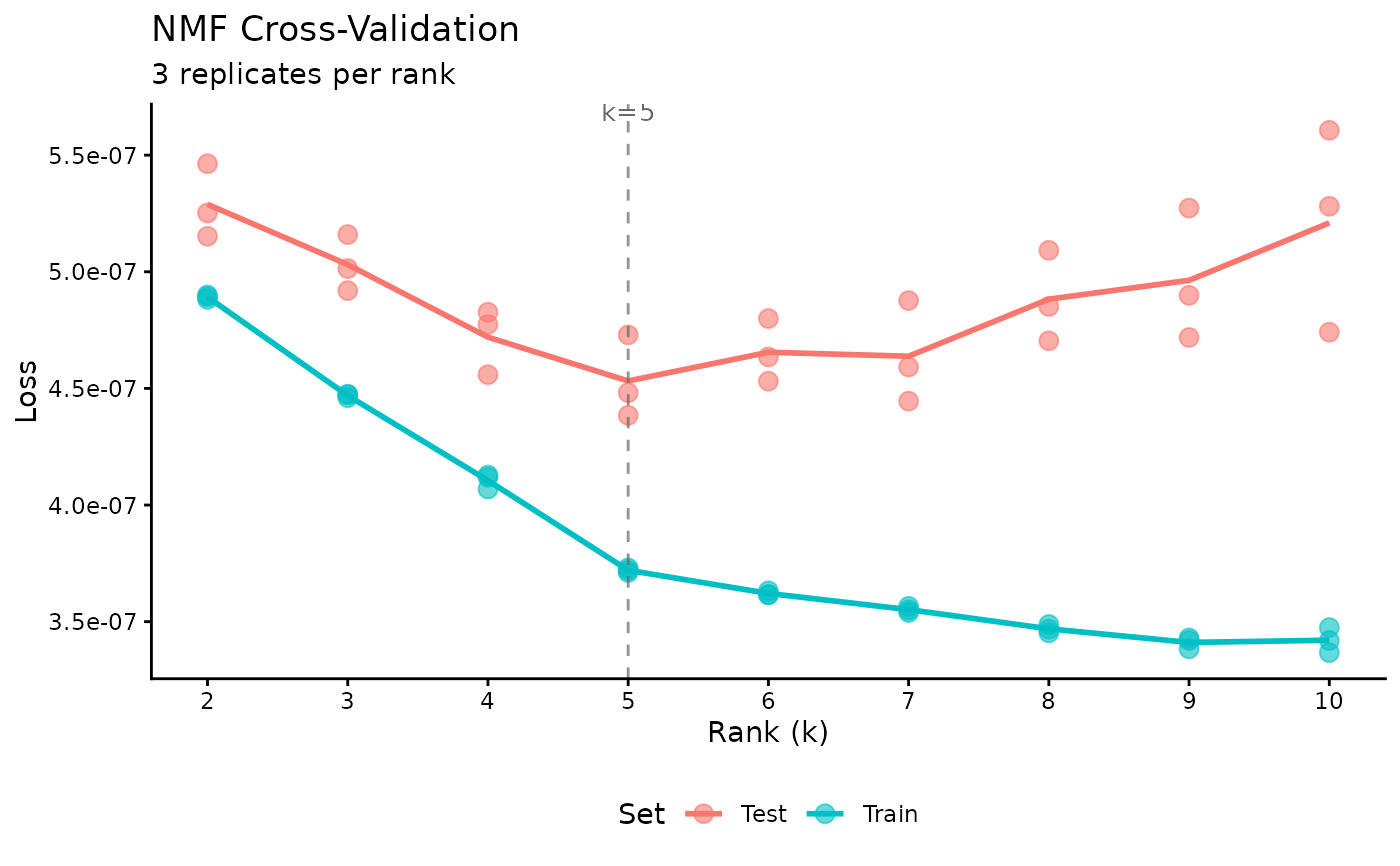 optimal_k <- cv$k[which.min(cv$test_mse)]
# fit final model with optimal rank
model <- nmf(sim$A, optimal_k)
# }
optimal_k <- cv$k[which.min(cv$test_mse)]
# fit final model with optimal rank
model <- nmf(sim$A, optimal_k)
# }